Version 2026.1 of the IUPHAR/BPS Guide to Pharmacology database was released on 12th March 2026, our first database release of the year. This blog post gives details of the key content updates and website changes. GtoPdb now contains:
- 3,127 human targets, 1,807 of which have curated quantitative ligand interactions.
- 13,721 ligands, 9,895 of which have curated quantitative target interactions.
- 2,196 approved drugs, 1,238 with curated quantitative interactions.
- Clinical use summaries for 4,257 ligands.
- 24,449 curated interactions, 22,189 which are quantitative.
- Data curated from 49,293 references
In this release, 130 new ligands, 132 new curated interactions have been added.
Commercial Access to GtoPdb
As a reminder to all our users, the Guide to Pharmacology has always been an open-access, freely available resource. Our expert-curated database of pharmacological data has been maintained over the past 10-15 years on ever changing and sadly, diminishing financial resources. We really want to keep GtoPdb open-access for all, but maintaining and updating the database is not free. Unfortunately the current funding landscape threatens our ability to ensure the Guide to Pharmacology’s sustainability going forward.
We have therefore taking steps to ask for commercial organisations, that use GtoPdb, to contribute financially to support its future maintenance, curation and sustainability.
More information can be found on our website: GtoPdb – Financial Sustainability Support
Curation Update
New Targets
New protein targets are added to the GtoPdb when evidence of their therapeutic potential is uncovered. Some of these require addition of new target families, whilst others are added to expand existing families. To complement these new targets we curate agents that have been shown to modulate target activity or functions, either as experimental probes and/or clinical leads.
The table below summarises the new targets that have been curated since our 2025.2 release (June 2025).
| TID | Family | Gene | Name | Comment |
| 3318 | Mitochondrial calcium uniporter (MCU) complex | MICU1 | mitochondrial calcium uptake 1 | Regulatory subunit of MCU; 2 experimental probe inhibitors curated |
| 3319 | Mitochondrial calcium uniporter (MCU) complex | MICU2 | mitochondrial calcium uptake 2 | Regulatory subunit of MCU |
| 3320 | Mitochondrial calcium uniporter (MCU) complex | MICU3 | mitochondrial calcium uptake 3 | Regulatory subunit of MCU |
| 3321 | Mitochondrial calcium uniporter (MCU) complex | MCU | mitochondrial calcium uniporter | Pore forming subunit of MCU |
| 3322 | Mitochondrial calcium uniporter (MCU) complex | MCUB | mitochondrial calcium uniporter dominant negative subunit beta | Pore forming subunit of MCU |
| 3323 | Mitochondrial calcium uniporter (MCU) complex | SMDT1 | EMRE (single-pass membrane protein with aspartate rich tail 1) | Pore forming subunit of MCU |
| 3324 | RAS subfamily | ARF6 | ARF GTPase 6 | Oncology target; inhibitor tool compound curated |
| 3325 | C19: Ubiquitin-specific protease | USP28 | ubiquitin specific peptidase 28 | Oncology target; inhibitor tool compounds curated |
| 3326 | Basic helix-loop-helix (BHLH) TFs | TCF4 | transcription factor 4 | Endogenous β-catenin binding protein, forming TF complex |
| 3327 | Zinc finger TFs | ZBTB7A | zinc finger and BTB domain containing 7A | Inhibition reactivates fetal hemoglobin (HbF) expression, for Sickle cell treatment; Ph1/2 molecular glue degrader curated |
| 3328 | RAS subfamily | TBC1D15 | TBC1 domain family member 15 | Involved in mitochondria-lysosome interactions and autophagy; experimental inhibitor probe curated |
| 3329 | Spliceosome proteins | RBM39 | RNA binding motif protein 39 | Oncology target; the aryl sulfonamides indisulam and tasisulam acts as molecular glue degraders |
| 3330 | Nucleoside synthesis and metabolism | SAMHD1 | SAM and HD domain containing deoxynucleoside triphosphate triphosphohydrolase 1 | Additional target of the EPHrin receptor inhibitor NVP-BHG712 |
| 3331 | Immunoglobulin like domain containing proteins | TREM2 | triggering receptor expressed on myeloid cells 2 | Functions as a lipid-sensing receptor; target for neurodegenerative diseases; agonist/activator tool compounds and Ph1 clinical candidate curated |
| 3332 | Basic helix-loop-helix (BHLH) TFs | OLIG2 | oligodendrocyte transcription factor 2 | CNS-specific TF; ongogene in some CNS tumours, oncology target; inhibitor CT-179 granted FDA orphan drug designation to treat gliomas (2017) |
| 3334 | 2-Acylglycerol ester turnover | ABHD4 | abhydrolase domain containing 4, N-acyl phospholipase B | Another enzyme in the family; no selective inhibitors reported |
| 3335 | Basic helix-loop-helix (BHLH) TFs | MYC | MYC proto-oncogene, bHLH transcription factor | Important oncogene; no direct small molecule inhibitors reported but a PROTAC has been curated; “Interdictor” molecules are being designed that interfere with MYC translation at the ribosome |
| 3336 | S1: Chymotrypsin | MASP2 | MBL associated serine protease 2 | Anti-MASP2 antibodies are proposed to reduce thrombotic microangiopathy and blood vessel damage |
| 3337 | Kinetochore proteins | SUGT1 | SGT1 assembly cochaperone of MIS12 kinetochore complex | Target of natural product Notoginsenoside-Fa; added in response to BJP author request for hyperlinks |
| 3338 | Rho GEFs | VAV1 | vav guanine nucleotide exchange factor 1 | Lymphoma oncology target; 2 molecular glue degraders curated, one is a Ph1 clinical candidate |
| 3339 | STAT transcription factors | STAT1 | signal transducer and activator of transcription 1 | Obesity-driven STAT1 signaling in the liver has been linked to metabolic dysfunction-associated steatohepatitis (MASH); experimental inhibitor probe curated |
| 3340 | Gelsolin/villin cytoskeleton proteins | AVIL | advillin | Oncogene in glioblastoma; small molecule inhibitor of binding to β-actin curated |
| 3341 | Methionine turnover | MAT2A | methionine adenosyltransferase 2A | Crucial for SAM biosynthesis in MTAP-deficient cancers, and proposed as synthetic lethal target; several allosteric inhibitors curated including one former clinical lead (Servier; development terminated in 2022) |
| 3342 | Macrophage migration inhibitory factor (MIF) family proteins | DDT | D-dopachrome tautomerase | Cytokine-like activity, implicated in tumorigenesis and inflammation; experimental inhibitor curated |
Ligands
Approved drugs 2025
Curation totalled 100 for all jurisdictions that we are able to track. FDA and EMA drug approvals are relatively easy to track (https://www.fda.gov/drugs/novel-drug-approvals-fda/novel-drug-approvals-2025, and download available from https://www.ema.europa.eu/en/medicines/download-medicine-data). The Nature Reviews Drug Discovery article covering approvals from the China NMPA was a very useful addition to the available literature for 2025 (https://pubmed.ncbi.nlm.nih.gov/41688808/).
Those from the 2025 set that have not been fully curated in the GtoPdb include antivirals, imaging reagents, enzyme and growth hormone replacement drugs and a few ADCs. Cell and gene therapies, and vaccines are not included in the GtoPdb.
The complete list is available here https://www.guidetopharmacology.org/GRAC/DrugApprovalsForward. We have update this table to include hyperlinks to both the ligand pages and to the primary therapeutic targets associated with the drugs.
FDA approved drugs 2026
There have been 5 FDA approvals so far in 2026:
- difamilast (Adquey), a PDE4D inhibitor to treat atopic dermatitis
- copper histidinate conjugate (Zycubo) as copper replacement for Menkes disease (defective ATP7A transporter-mediated copper absorption)
- milsaperidone (Bysanti), an iloperidone prodrug for schizophrenia
- pegzilarginase-nbln (Loargys), ERT to treat hyperarginemia in arginase 1 deficiency
- navepegritide (Yuviwel), a growth hormone analogue to treat achondroplasia
WHO INN proposed list 134 (Feb 2026)
There are 202 INNs for human medicaments in the list, with 172 that are not cell/gene therapies, vaccine agents, enzyme replacement therapy, or radiodiagnostic agents. There were 2 replacement INNs for those in previous lists. There is a brief COVID-19 (special edition) included which lists just one spike protein-based agent for anti-SARS-CoV-2 immunization.
The table below shows the 36 INNs from list 134 that are curated in the GtoPdb. The 12 with LIDs <14180 were already curated and for these the new INNs have replaced the existing ligand names. The remaining 24 are newly curated ligands. We have made progress in associating the INNs with potential company research codes, pharmacological data (generally from patents) and clinical trial information. This work will continue when staffing resources permit.
| LID | Name | Comment |
| 5290 | ninvosudil | Rho kinase inhibitor |
| 7000 | curcumin | antioxidant |
| 8741 | resveratrol | antioxidant |
| 12300 | lemuplamor | epoxide hydrolase 2 inhibitor; thrombolytic for acute ischemic stroke |
| 12892 | tipasertideg | IRAK4 degrader; Ph1 atopic dermatitis (AD) or hidradenitis suppurativa |
| 13004 | elticertinib | Mutant EGFR inhibitor; Ph1 Ex20 insertion +ve NSCLC |
| 13018 | romoverlib | WRN RecQ like helicase inhibitor; Ph1 solid tumours |
| 13132 | sorfequiline | Antibacterial; Ph2 TB |
| 13273 | pexerelgon | RXFP1 agonist; Ph1 chronic heart failure |
| 13919 | clifutinib | FLT3 RTK inhibitor; Ph3 FLT3-ITD AML |
| 14175 | naperiglipron | small molecule GLP1R agonist; Ph2 obesity or overweight |
| 14179 | canverixin | ACKR3 (CXCR7) antagonist immunomodulator; Ph1 healthy subjects |
| 14354 | embibatinib | ABL kinase inhibitor; Ph1/2 CML |
| 14355 | fimsosertib | HPK1 (MAP4K1) inhibitor; Ph1/2 advanced solid tumours |
| 14356 | liradasertib | LRRK2 inhibitor; Ph2 LRRK2-associated Parkinson’s disease |
| 14357 | meribatinib | ABL kinase inhibitor; Ph2 CML |
| 14358 | conveglipron | small molecule GLP1R agonist; preclinical |
| 14359 | mitucapistat | calpain inhibitor; Ph1 healthy subjects |
| 14360 | nomelcitinib | TYK2 inhibitor; Ph3 plaque psoriasis |
| 14362 | pelerasertib | LRRK2 inhibitor; Ph1 healthy subjects |
| 14363 | relzasertideg | casein kinase 1α degrader; Ph1 advanced AML or higher-risk myelodysplastic syndromes |
| 14364 | risimtinib | FGFR RTK inhibitor; development stage undetermined |
| 14365 | vanoglipel | GPR119 agonist; Ph2 MASH/NASH |
| 14366 | vepugratinib | FGFR RTK inhibitor; Ph1 solid tumours with FGFR mutations |
| 14367 | vexicitinib | JAK/SYK inhibitor; Ph3 plaque psoriasis |
| 14368 | vustanaciclib | CDK4/2 inhibitor; Ph1/2 breast cancer |
| 14369 | zinilisib | PI3Kα inhibitor; Ph1 solid tumours |
| 14370 | zecontimod | CD3-Nck PPI inhibitor; Ph1/2 plaque psoriasis |
| 14371 | relcobatinib | ABL1 allosteric inhibitor; Ph1 CML |
| 14372 | soralimixin | Antibacterial; Ph1 multidrug-resistant (MDR) Gram-negative bacterial infections |
| 14373 | dezandrodeg | AR PROTAC; Ph3 met castration-resistant prostate cancer |
| 14377 | forazapadin | AAK1 inhibitor; Ph2 DMD |
| 14378 | balumorexton | OX2 receptor agonist; Ph2 sleep disorders |
| 14380 | gosicalcetide | CaSR agonist; secondary hyperparathyroidism |
| 14415 | parunoflast | CNS-penetrant NLRP3 inhibitor; Ph2 Parkinson’s, obesity-associated chronic Inflammation |
| 14416 | abdenoflast | Peripherally active NLRP3 inhibitor; Ph2 pericarditis, cryopyrin-associated periodic syndrome |
Antibacterial Curation
Our collaboration with Antibiotic DB (ADB; www.antibioticdb.com) continues to allow us to extend the coverage of ligands with annotated antibacterial activity in GtoPdb and provide comprehensive chemistry and pharmacology for select antibacterials curated within ADB, via reciprocal links. This project is supported by the Global Antibiotic Research and Development Partnership (GARDP; https://gardp.org/).
Currently we have 738 ligands tagged in GtoPdb as ‘antibacterial’ and 716 of these have links to compounds at ADB. 276 are approved drugs. Since our last release, the number of antibacterial tagged compounds in GtoPdb has increased by 25, with 22 of these being newly-added ligands.
We are also please to see the publication of the following paper, which gives an excellent overview of updates to the AntibioticDB resource:
Galarion LH, Hennessy A, Harding SD, Armstrong JF, Pentz-Murr A, Davies JA, O’Neill AJ, Piddock LJV. AntibioticDB: An Updated and Improved Open-Access Database for the Antibacterial Research and Development Community. ACS Infect Dis. 2026 Feb 23. doi: 10.1021/acsinfecdis.5c00955. Epub ahead of print. [PMID: 39146948]
See also our blog post on this publication.
Website Updates
Downloads and web-service update
GtoPdb Downloads: https://www.guidetopharmacology.org/download.jsp
Additional reference columns have been added to our interaction download files. Previously these files only listed references that have PubMed IDs. This has now been extended to include columns for references that are web-pages or patent numbers. We are looking to extend this to all journal references, irrespective of whether they have PubMed IDs.
Updated sub-structure search
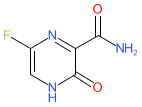
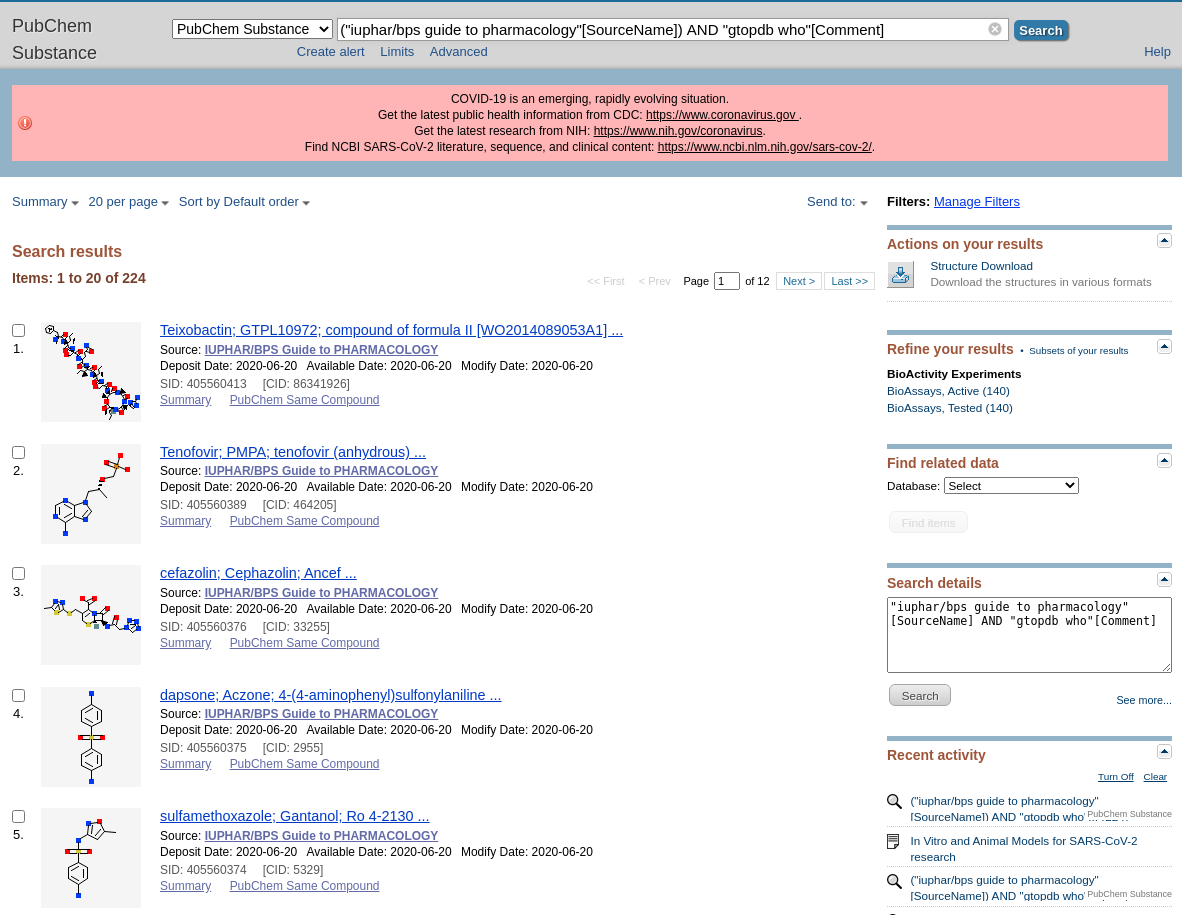
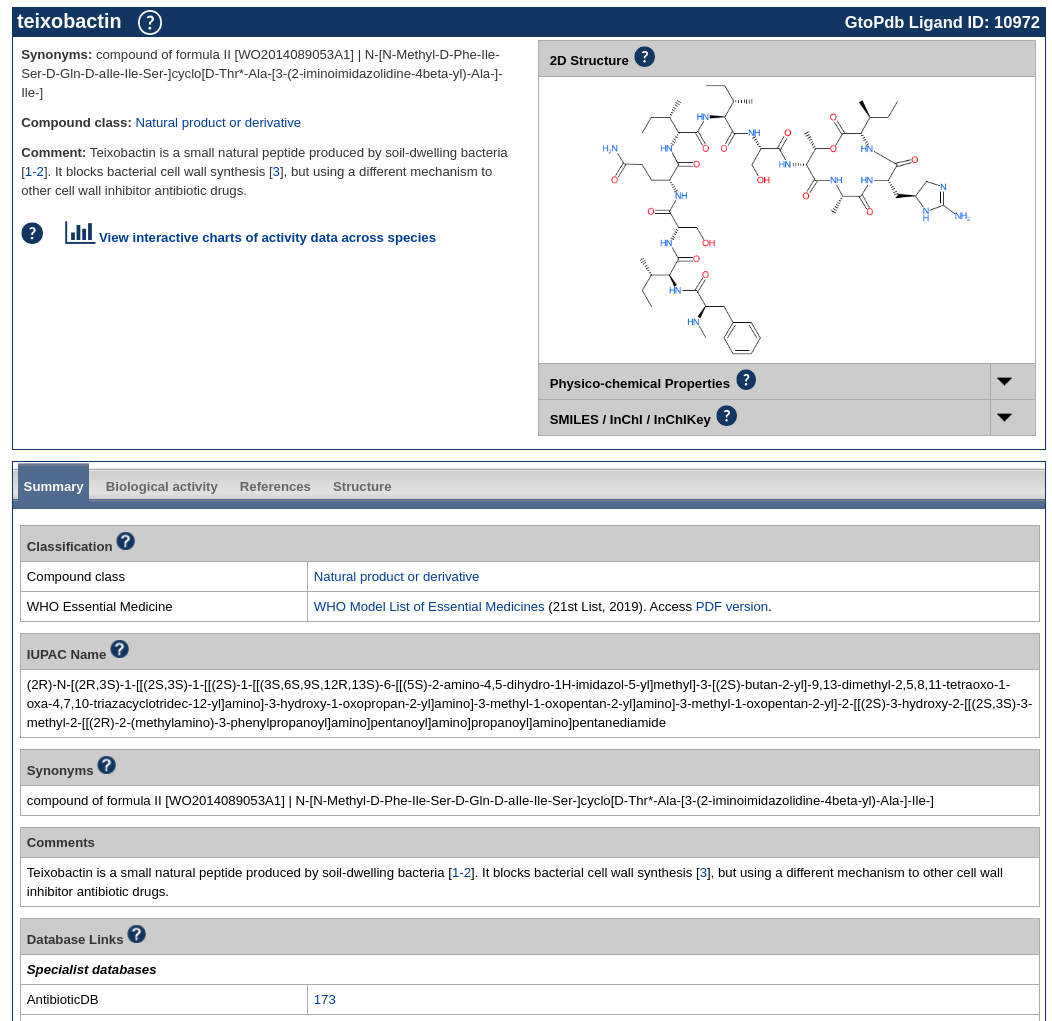
A reminder that we recently updated our chemical structure search to include highlighting the query structure in result sets from a substructure search. This uses Chemistry Development Kit’s (CDK) depict feature and highlight functionality. When a user runs a chemical structure search and sets the type to substructure, the query structure will be highlighted in the results set.
The below shows an example of a result set from a substructure search. Showing the query structure highlighted (in cyan).

In addition to adding highlighting on substructure searches, we are also implementing the display of the Tanimoto Coefficient value used when a similarity search is run.
Drug Approvals summary page
As mentioned earlier we have made some minor updates to our drug approvals page https://www.guidetopharmacology.org/GRAC/DrugApprovalsForward, to include hyperlinks to the primary therapeutic targets associated with the drugs






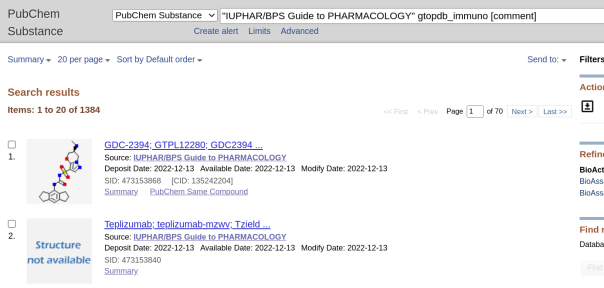









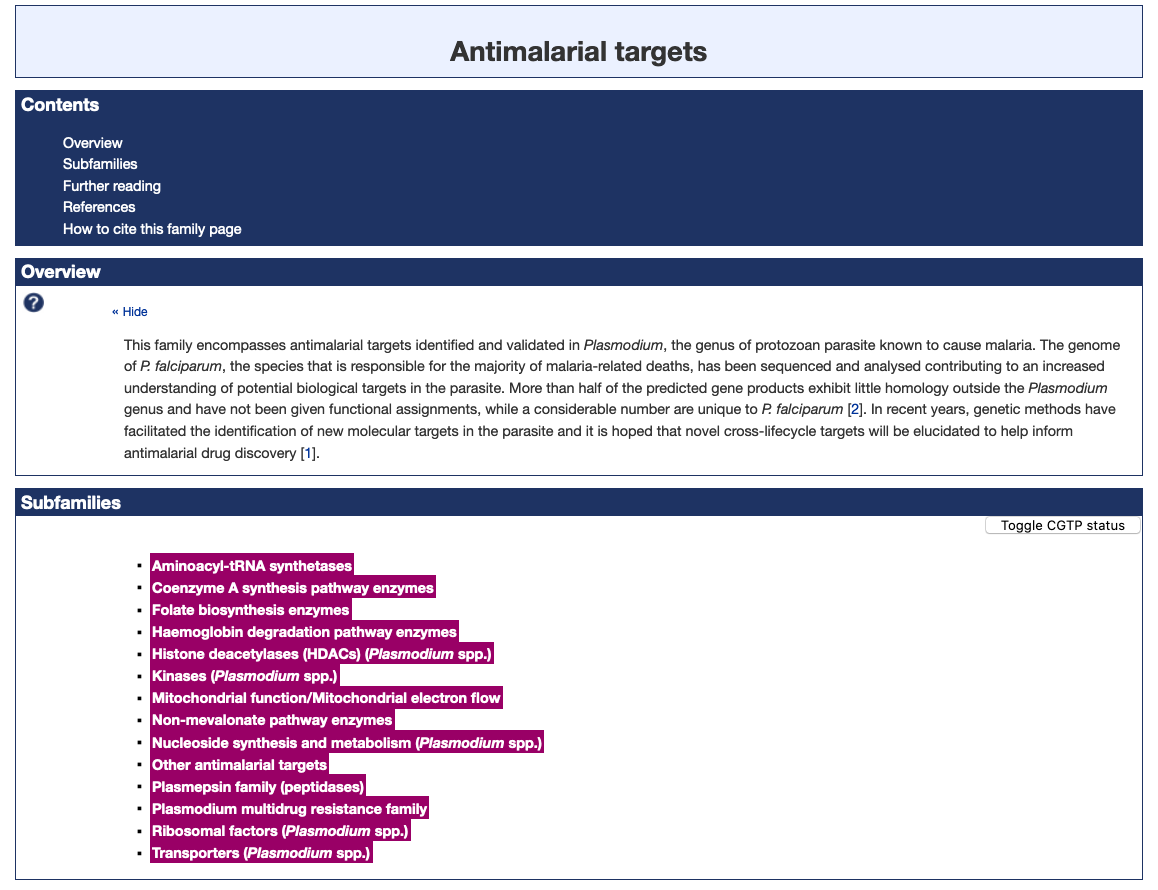 Figure 1. The Antimalarial targets family page illustrating the new subfamily classification (highlighted in magenta).
Figure 1. The Antimalarial targets family page illustrating the new subfamily classification (highlighted in magenta).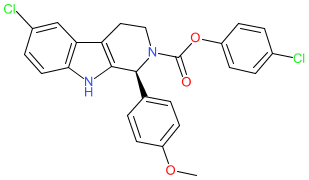




 In
In