This post on SID tagging has been reproduced, with permission, from Dr. Chris Southan’s original post in his blog – Bio <-> Chem.
It is intended to be user-orientated, those interested in the technicalities are welcome to contact the Guide to Pharmacology curation team. The links and counts in this post we taken on 19th Dec 2022 and reflect data from the 2022.4 GtoPdb release.
Looking at the tags we have introduced will make this clearer. For starters, as many may know, as of release 2022.4 GtoPdb has 11603 PubChem substance (SID) records. Looking at SID472319339 we can find the following comments.
This includes the explicit tags
gtopdb_approved – Substance is an approved drug in GtoPdb.
gtopdb_antibacterial – Substance is tagged as an antibacterial in GtoPdb.
So we can employ an interface selection for the gtopdb_approved tag to get 1813 “approved” (you don’t usually need the [comment] as a field restrict but useful for more complex queries).
Similarly we can select as per below for the 365 antibacterial
We can then make the requisite intersect via the “Advanced” option on the query interface as below:
The intersection search is thus (“IUPHAR/BPS Guide to PHARMACOLOGY”[All Fields] AND gtopdb_antibacterial[comment]) AND (“IUPHAR/BPS Guide to PHARMACOLOGY”[All Fields] AND gtopdb_approved[comment]) with the result of 146
We have introduced an additional three selects. The first is
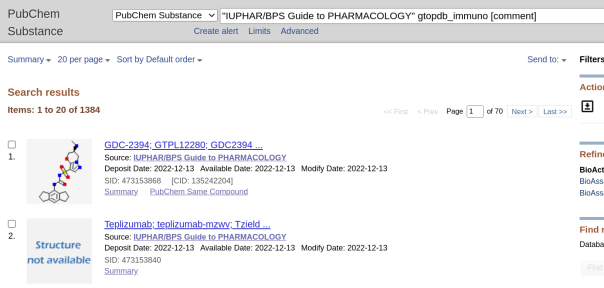
gtopdb_immuno – Substance is curated in IUPHAR Guide to Immunopharmacology (GtoImmuPdb)
with 1384 SIDs as per below:
Cutting the story short we can easily spot the difference between the small-molecule entries that will be subsumed into Compound (CIDs) and the “Structure not Available” entries (two above) that, as large macromolecules, cannot form CIDs and will thus remain SID-only entries
These are selectable via our fourth tag
gtopdb_antibody – GtoPdb identifies this substance as an Antibody
With which we can select the 335 antibody entries below :
As performed above the intersect below gives the 127 approved antibodies:
Last but not least our fifth tag is
gtopdb_malaria – Substance is curated in IUPHAR/MMV Guide to Malaria Pharmacology (GtoMPdb)
Combining with Venny
Either by combining queries or (much easier) using the query history under the advanced tag we can make any sequence of Boolean selects (i.e. AND, OR, NOT) from combinations of our five tags for “approved” “immuno” “antibiotic” “antibody” or “antimalarial”. Below we can see another way of slicing and dicing via Venny. The example result shown below compares three of the tag lists we have already looked at.

Another advantage of these GtoPdb specific retrievals is to be able to pivot across to small molecule CIDs. The steps to do this are shown below, highlighted in yellow.
The result below, transformed into a PubChem compound query, now retrieves all CIDs that include GtoPdb approved SIDs
Thus GtoPdb contains 1586 approved drug CIDs. Note that not all these are strictly “small molecules as we can see from the Mw ranking below where the top-ten are above 5000 Mw













Leave a comment