The latest release of the Guide to PHARMACOLOGY database, version 2020.3, has now been made. A large focus of our curation over the last couple of month since our 2020.2 release We are pleased to have been able to make an expeditious database release (version 2020.2), following on from our last update in March 2020 in order to make public new curation specifically related to SARS-CoV-2.
Coronavirus
The Guide to Pharmacology coronavirus information page continues to be updated on a regular basis to capture the latest pharmacological strategies under investigation to mitigate against COVID-19.
Recently published in BJP is our review that “sets out to identify opportunities for drug discovery in the treatment of COVID‐19 and in so doing, provide a rational roadmap whereby pharmacology and pharmacologists can mitigate against the global pandemic“:
Alexander SPH, Armstrong J, Davenport AP, Davies JA, Faccenda E, Harding SD, Levi-Schaffer F, Maguire JJ, Pawson AJ, Southan C, Spedding MJ. (2020). A rational roadmap for SARS‐CoV‐2/COVID‐19 pharmacotherapeutic research and development. IUPHAR Review 29 Br J Pharmacol. doi: 10.1111/bph.15094. [PMID:32358833]
Content Updates
Main target families with updated in this release:
GPCRs
- Chemokine receptors
- Corticotropin-releasing factor receptors
- Dopamine receptors
- Formylpeptide receptors
- Glucagon receptor family
- Histamine receptors
- Lysophospholipid (S1P) receptors
- Opioid receptors
- Proteinase-activated receptors
- Urotensin receptor
Ion Channels
NHRs
Enzymes, Catalytic Receptors
- Akt (Protein kinase B, PKB) family
- Beta-adrenergic receptor kinases (âARKs)
- M10: Matrix metallopeptidase
- N-Acylethanolamine turnover; in particular N-Acylphosphatidylethanolamine-phospholipase D has had 4 inhibitor compounds added
- RAS subfamily
- Toll-like receptor family
New drug target
Acyl-CoA synthetase short chain family member 2 (ACSS2) is an emerging oncology drug target that plays a role in epithelial-mesenchymal transition (EMT) and cancer cell metastasis. Two ACSS2 inhibitor probe compounds have been curated.
AntibioticDB
As part of a collaboration with AntibioticDB (https://www.antibioticdb.com/), we have now begun to identify and tag sets of antibiotic ligands in GtoPdb. Where we have identified mappings between these and compounds in the AntibioticDB repository, we’ve put in place direct links. The newly curated set includes approved and investigational antibiotics and pre-clinical leads, as well as compounds whose development has been discontinued.
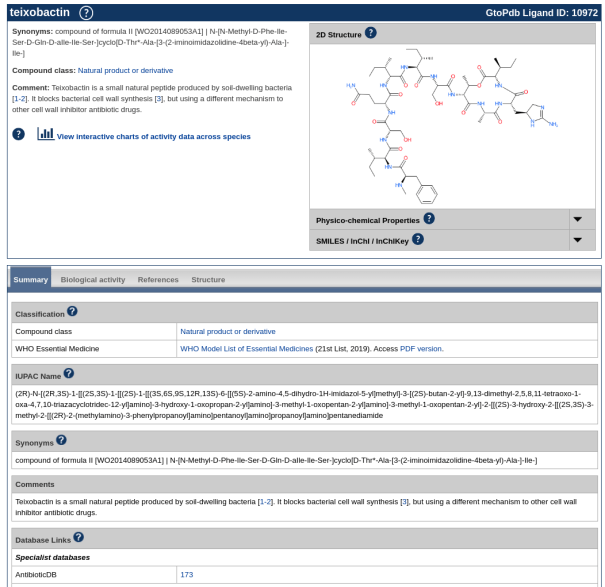
Teixobactin ligand summary page (https://www.guidetopharmacology.org/GRAC/LigandDisplayForward?ligandId=10972). At the bottom, a new specialist database link now existing to the mapped entry in AntibioticDB.
Website Updates
Ligand Summary Pages
The layout of the ligand summary pages has been changed to hopefully provide more of the key information on a ligand at the top of these pages. In the example below we show how this looks for chloroquine.
The main information box now contains:
- Synonyms
- Icons to indicate key ligand classifications
- Drug approval indication
- GtoPdb curator comments
- Link to ligand activity graphs
In addition, the 2D ligand structure is display beside this information in a expandable/retractable section where users can also view physico-chemical properties and SMILES/InChI/InChI Keys.
Previously the top part of the ligand summary pages contained too much white space, repetitions of the ligand name and emphasised the ligand ID, physico-chemical properties and to some extent buried comments, SMILES/InChI keys and useful links (such as to the ligand activity graphs).
The reorganisation of the page now emphasises key information. GtoPdb curator comments in particular contain a valuable description of the ligand and provide explanations as to why they have been curated in GtoPdb.
WHO essential medicines
Any ligands included in the World Health Organization (WHO) Model List of Essential Medicines (21st list, 2019) are now tagged in GtoPdb. This means users can easily view these ligands on our ligand list page:
https://www.guidetopharmacology.org/GRAC/LigandListForward?type=WHO-essential&database=all
It is also possible, as it is for all ligand lists, to download this set from this page by clicking the download link in the top-right of the table (see below).

Other Updates
ChEMBL 27
Both the ligand activity graphs and pharmacology search tools have now been updated to access the latest ChEMBL release (ChEMBL 27).
Links to DrugCentral 2020
We have added links to the DrugCentral 2020 Online Drug Compendium. DrugCentral provides information on active ingredients chemical entities, pharmaceutical products, drug mode of action, indications, pharmacologic action. It monitors FDA, EMA, and PMDA for new drug approval on regular basis to ensure currency of the resource. Limited information on discontinued and drugs approved outside US is also available however regulatory approval information can’t be verified. The database is developed and maintained by Oleg Ursu and Tudor Oprea and the web application is developed by Jayme Holmes.
We have so far mapped 1433 ligands in GtoPdb to entries in DrugCentral.



Leave a comment