A hallmark of GtoPdb is our curation of out-links as opposed to adding these by automated cross-referencing. The latter can give rise to not only false +ves and false -ves but also 1:many relationships that users find difficult to resolve. Also valuable are in-links from other relevant resources. Not only do these facilitate reciprocal navigation but also linked-data queries via inter-database web services. The Edinburgh team engages extensively with other databases at many levels, including long-standing collaborations and conference catch-ups. Indeed, a component of our value is expert selection of out-links for our GtoPdb entries, especially since pharmacology spans the domain complexity of bioinformatics, chemistry and genomics. Reciprocity of linking (a.k.a. cross-pointing) between any two databases thus becomes an enabling feature for both.
This document reviews in-links to GtoPdb from public sources that have come to our attention. However, there may be others we are not aware of (e.g. inside pharmaceutical companies). Those resources that we specifically out-link to are listed in Table 5 in our NAR article PMID 26464438. There are other cases where reciprocal in-links are under consideration but not yet instantiated (e.g. NURSA , ESTHER DrugBank and Open PHACTS).
As an open database we welcome relevant in-links. Notwithstanding, there are caveats, especially where these have been instigated without contact or our technical input. Two main problems arise. Firstly automated parsing may not be consolidated by specificity checks on their side. Secondly their update frequency my not be synchronised with our own new releases. To assess the latter we can use source entity counts and loading dates but these are not always provided. For example, we have been contacting resources we know who have not yet replaced IUPHAR-DB content by GtoPdb but its difficult to find all instances. Those in-links we know of are listed below (but please contact us if you are aware of others).
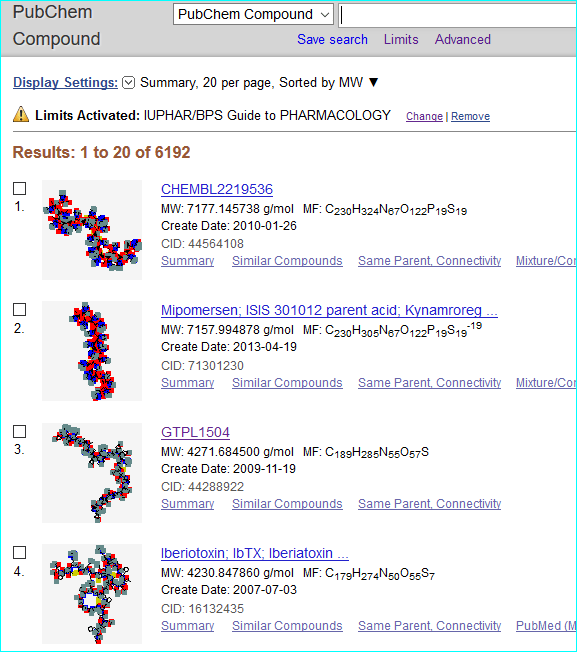
For PubChem (a global chemistry and bioactivity portal) we have 8201 ligand submissions as SIDs that each include a GtoPdb url. Of these 6192 are merged into Compound identifiers (CIDs) with a defined chemical structure. Most of the SIDs 2009 SIDs without CIDs are large peptides, small proteins and antibodies that cannot form a CID. Note we have some SID duplicates structures where we have separate GtoPdb ligand IDs pointing to radiolabel citation data without specified substitution positions that have a CID. We also have 55 BioAssay entries for 5HT sub-family chemistry mappings.
HGNC is responsible for approving unique symbols and names for human loci, including protein coding genes, ncRNA genes and pseudogenes. We have a long-standing collaboration via NC-IUPHAR. An example outlink is shown below.
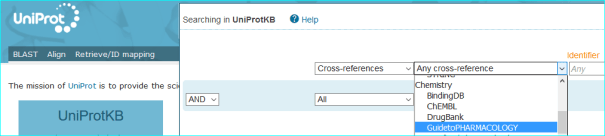
UniProtKB/Swiss-Prot: We are included in the Cross-References for protein entries. These can be selected using the menu below
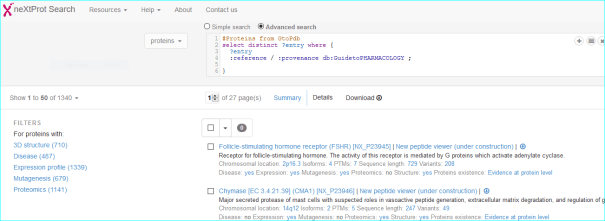
The query currently produces 1829 proteins with GtoPdb links as having quantitative ligand interactions.
neXtProt (PMID 25593349) is a protein-centric knowledgebase developed at the SIB Swiss Institute of Bioinformatics focused solely on human proteins. In a sense this is “forked” from Swiss-Prot but is technically distinct. It inherits our UniProt links.
IMGT/mAb-DB is a high-quality integrated information system focused on clinical antibodies. We have a long-standing collaboration. A link example is shown below.

ChEMBL, a database of bioactive drug-like small molecules with calculated properties and abstracted bioactivities. A target link example is below.
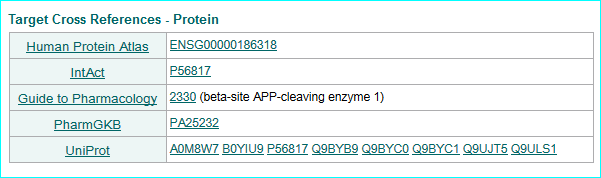
Our ligands get a nested link in the ChEMBL interface via UniChem.
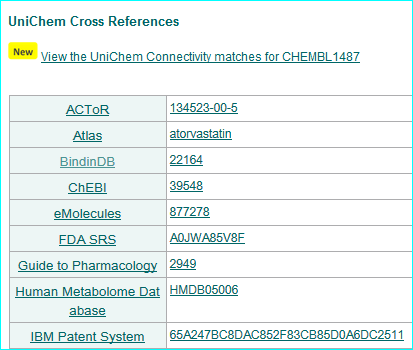

UniChem produces cross-references between chemical structure identifiers from different databases within the EBI. We are listed as a source

This was updated on 17-NOV-15 as 6006 chemical entities
***********************************************************************
MEROPS information resource for peptidases (also termed proteases, proteinases and proteolytic enzymes) and the proteins that inhibit them. An example of our cross-links is shown below.
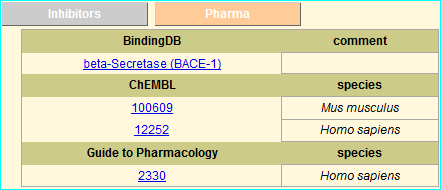
The addition of these links is specified in the latest MEROPS NAR publication (PMID 26527717)
GPCRdb (in new NAR as PMID 26582914) , the information system for G protein-coupled receptors. Consequent to extensive contact and collaboration we have mentions in their paper and web resource. In addition we have pioneered reciprocal web services. A link example is shown below.
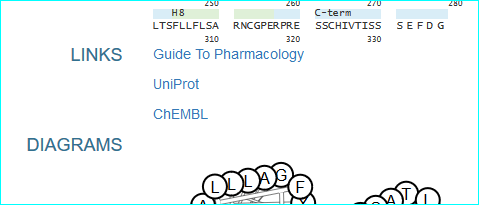
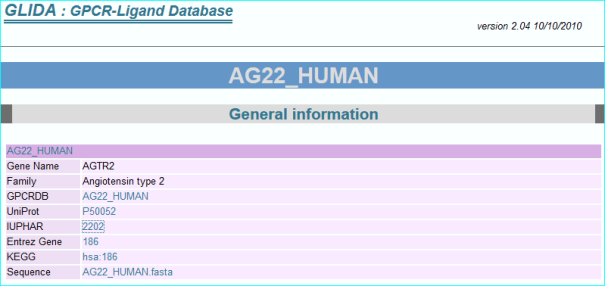
ChemSpider. This is a leading chemistry portal of 34 million compounds and also a reference structure source for OpenPhacts
We can see from this example link for CS19973960 that we have IDs but no direct outlink at the moment.
***********************************************************************
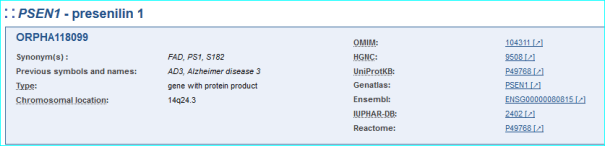
Oprhanet is the portal for rare diseases and drugs with whom we have regular contact. We are one of the eight gene-mapped IDs (see Part IV in the user guide and sample entry below)
CARLSBAD is an integrated resource based on filtered subsets from the bioactivity databases listed below (PMID 23794735).
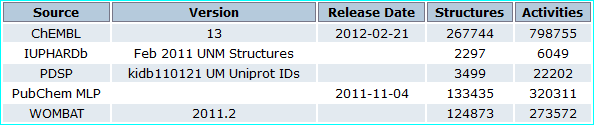
Thus the 2012 release subsumed the 2011 IUPHAR-DB set of 2297 ligands.
In the GLASS database of GPCR-ligand associations (PMID 25971743) we are cited as one their five sources. An example x-ref is shown below but the statistics are not specified.
ChemProt -2.0 (PMID 23185041).
This resource with an emphasis on visual navigation in a disease chemical biology specifies eight protein < > ligand sources collated in 2012. X-ref counts were not specifies but this includes IUPHAR-DB from 2011.
The RefSeq LinkOut feature facilitates access to relevant online resources beyond the NCBI Entrez system. These include GtoPdb in the protein section with the example for NP_036236 (BACE1) shown below.

ZINC A free database of commercially-available compounds for virtual screening (PMID 26479676). The x-ref below is for IUPHAR-DB
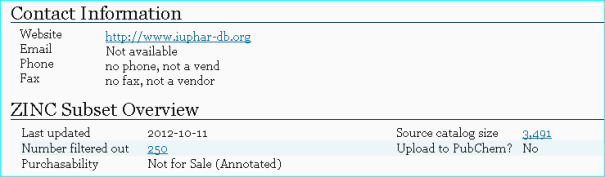
ZINC has just undergone a major update in PubChem so our entries can be intersected
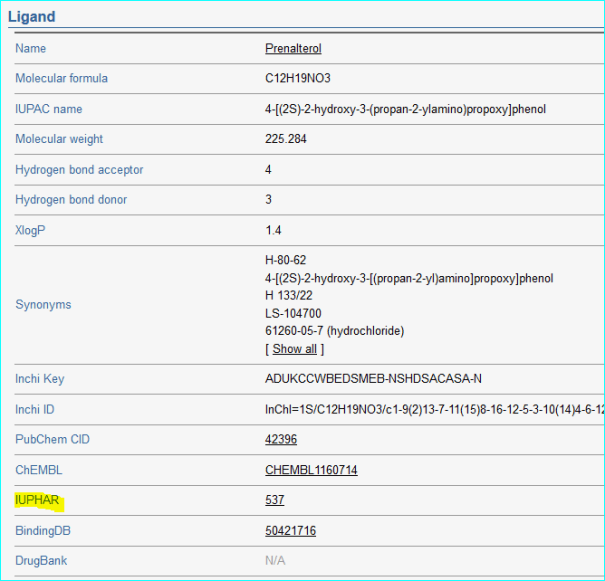
GeneCards automatically integrates gene-centric data from ~125 web sources. As we can see we became source 104 (but named IUPHAR)

We have had collaborative contact w.r.t. to their MalaCards disease database where we are source 28.

DGIdb 2.0 is a resource for mining clinically relevant drug-gene interactions (PMID 26531824). They quote on their paper “new content was imported from the IUPHAR/BPS Guide To Pharmacology (GTP) (6), accounting for 10 225 interaction claims and 1,969 druggable gene category claims” . An example of a search result specific for us, is shown below
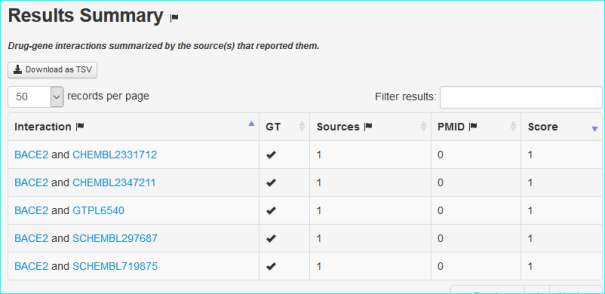
We also get a chemistry page, with the download date
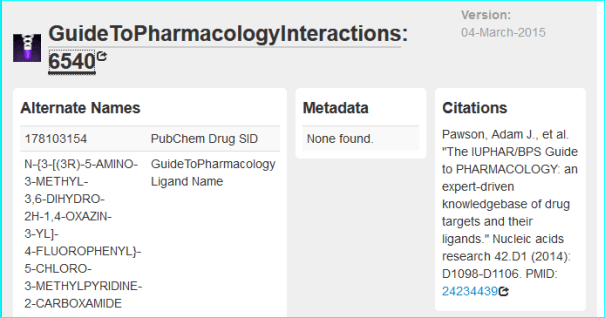
And a gene page (below)
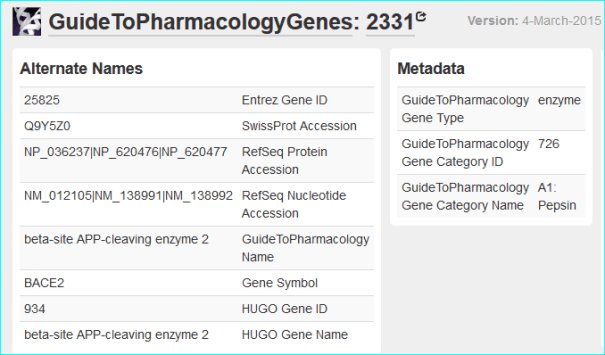
They have initiated contact and cross-mapping issues can now be addressed directly via GitHub.
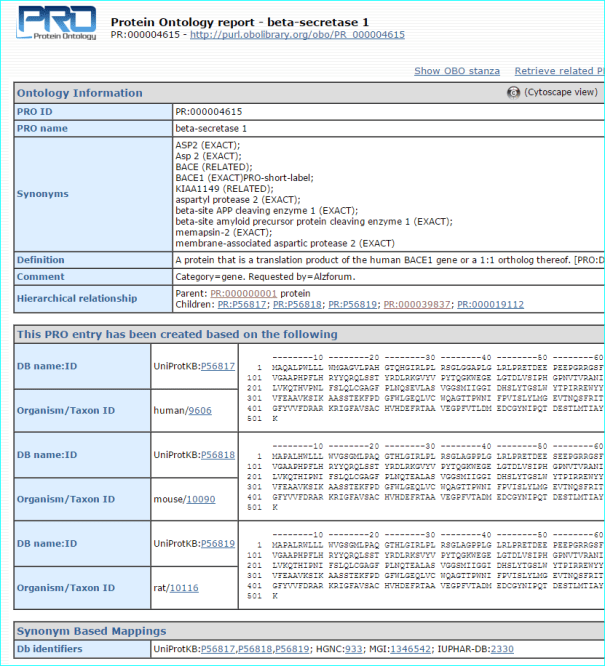
We are indexed in the Protein Ontology resource that provides an ontological representation of protein-related entities by explicitly defining and showing the relationships between them.
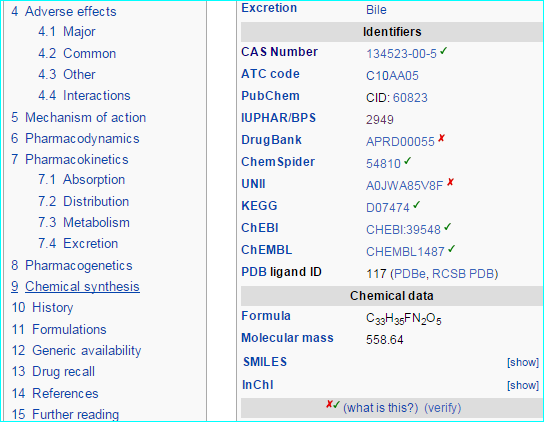
Wikipedians have been pointing to us for some time. Examples of an established target and a more recent ligand entry are shown below.



[…] of using a log-in but we do track usage and downloads for our own impact assessment. As already reported we were pleasantly surprised to discover various subsumations have spawned in-links we were unaware […]
[…] as part of integration efforts or simply bringing it inside their firewalls. When we looked at in-links (i.e. resources pointing to us) last year were surprised to find well over 20 of these. From […]