 We are pleased to make public the first beta-release (v1.0) of the Guide to MALARIA PHARMACOLOGY (GtoMPdb), a new extension to the existing Guide to PHARMACOLOGY (GtoPdb). The GtoMPdb is being developed as a joint initiative between Medicines for Malaria Venture (MMV) and the International Union of Basic and Clinical Pharmacology (IUPHAR), with the aim of adding curated antimalarial data to GtoPdb and providing a purpose-built portal that is optimized for the malaria research community.
We are pleased to make public the first beta-release (v1.0) of the Guide to MALARIA PHARMACOLOGY (GtoMPdb), a new extension to the existing Guide to PHARMACOLOGY (GtoPdb). The GtoMPdb is being developed as a joint initiative between Medicines for Malaria Venture (MMV) and the International Union of Basic and Clinical Pharmacology (IUPHAR), with the aim of adding curated antimalarial data to GtoPdb and providing a purpose-built portal that is optimized for the malaria research community.
We have implemented a number of changes to the existing database structure and web interface that were necessary for the capture and presentation of antimalarial data. Many antimalarial compounds have a poorly understood mechanism of action and an unknown molecular target and we have extended the interactions table and updated the web interface to accommodate this. A new “whole organism” assay type has been introduced to capture data from the whole cell assays used routinely in antimalarial drug discovery. Both changes are illustrated below.
Figure 1: The interactions table on an antimalarial ligand summary page
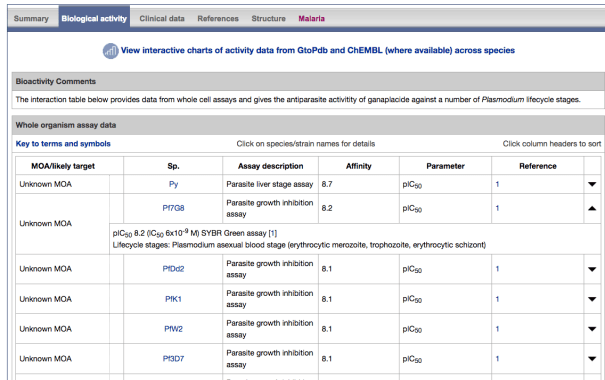
In addition, we have put in place the ability to tag both targets and ligands of relevance to malaria and provide curatorial comments. These comments surface on the website (see Figure 2 below) and are incorporated into the site search.
Figure 2: Malaria comments tab on an antimalarial ligand summary page
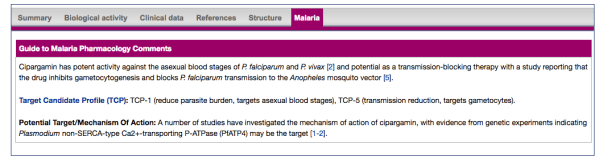

A new GtoMPdb portal (www.guidetomalariapharmacology.org) is being developed to provide tailored routes into browsing the antimalarial data in addition to the existing ligand and target browse/search functionality available on the parent GtoPdb site. For beta-release v1.0 we have implemented customised views of the data that include parasite lifecycle and target species activity, with access from either the menu-bar or panels on the homepage (see figure 3 below).
Figure 3: The GtoMPdb portal homepage

The GtoMPdb uses a set of top-level Plasmodium lifecycle stages (collective categories for one or more developmental forms of the parasite) against which interactions in the database can be annotated and which form the basis of organising, navigating and searching for parasitic lifecycle activity. We have developed a new Parasite Lifecycle homepage that provides a short introduction and links to additional pages for each of the top-level lifecycle stages. These in turn contain a more detailed description and a table of interactions for that lifecycle stage (illustrated in Figure 4).
Figure 4: Plasmodium liver stage page, an example of the new Parasite Lifecycle Stage pages

The Target Species homepage provides a short description for Plasmodium species that are of clinical or research importance. It also includes a resource section and links to individual pages for species that have annotated interactions in the database. The figure below illustrates an example of an individual species page. The interactions table displays affinity data for the species but also provides additional details, when available, for the strain used.
Figure 5: Plasmodium falciparum page, an example of the new Target Species pages

Development of the beta-release will continue with regular updates planned over the next few months as the quantity of data captured increases and improvements in the site layout and function are made.
If you have any feedback or queries about the resource please contact enquiries@guidetopharmacology.org
This project is supported by a grant awarded to Professor Jamie Davies at the University of Edinburgh by Medicines for Malaria Venture (MMV).


[…] this month we issue a blog post introducing the Guide to Malaria Pharmacology. This gave a very good background to the project and illustrated how we plan to handle curation of […]
[…] blog post gives more detailed information about the development of […]
[…] this year we issue a blog post introducing the Guide to Malaria Pharmacology. This gives a good background to the project and illustrates how we plan to handle curation of this […]