This blog-post will discuss the major developments planned for the Guide to IMMUNOPHARMACOLOGY as we look ahead to our next beta-release (v3.0) in early 2018.
This month, the updated IUPHAR/BPS Guide to PHARMACOLOGY NAR Database Issue has been published online (https://academic.oup.com/nar/article/4628131). [PMID: 29149325]
The IUPHAR/BPS Guide to PHARMACOLOGY in 2018: updates and expansion to encompass the new guide to IMMUNOPHARMACOLOGY
Nucleic Acids Research, gkx1121, https://doi.org/10.1093/nar/gkx1121
As the title indicates, a major part of this update includes the expansion of the database and developments to produce the new Guide to IMMUNOPHARMACOLOGY. The paper discusses the unique targets and ligands that have been incorporated into GtoPdb as a consequences of the GtoImmuPdb Project. For example targets of relevance to immunity, inflammation and infection such as pattern recognition receptors and protein of the innate immune response. Database content statistics are presented with a specific breakdown for GtoImmuPdb content (Table 1).
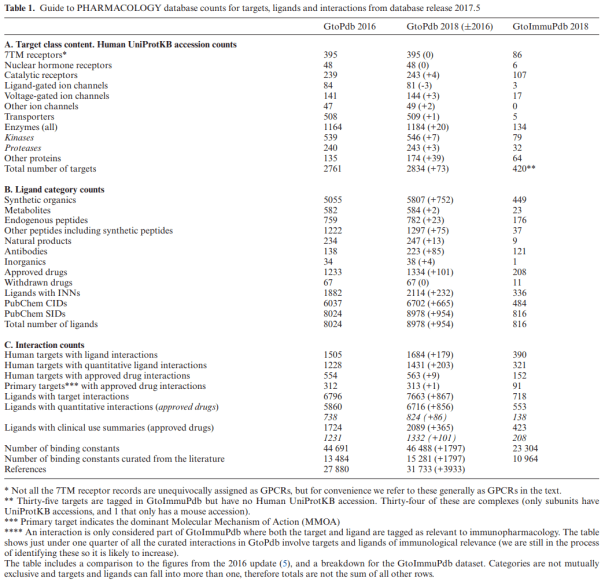
Table 1. Taken from the NAR paper, table gives a breakdown of database content statistics, including GtoImmuPdb counts.
The paper goes into details on the development of the Guide to IMMUNOPHARMACOLOGY in terms of content & curation and how targets and ligands of immunological relevance are identified. There is detailed discussion on the process of incorporating the new process, cell type and disease data types for GtoImmuPdb as well as explanations of the novel portal and interfaces that have been developed to surface the GtoImmuPdb data.
The discussion and descriptions in the paper are related to the beta-release v2.0. Our next planned beta-release is due in early 2018. The developmental priorities for this release are;
- Improving disease associations and display
- Graphical browsing / navigation
- Advanced search tool for immuno data types
- Video help tutorials
For the disease data we are looking at developing new disease summary pages. These will not only serve to display target and ligand associations to disease of immunological relevance – but will also capture and display all disease-related data in the GtoPdb. This includes pathophysiology data and information on mutations. We are currently working-up some prototype pages, but expect to be able to have have some form of disease pages available in beta v3.0.
Using graphical illustrations of key biological pathways and cell types, as a way to summarise data can be very valuable. Enabling such graphics to be interactive and support navigation of a website may bring added value to the GtoImmuPdb resource. We are at the early stages of developing a cell types graphical-based navigation tool (Figure 2).
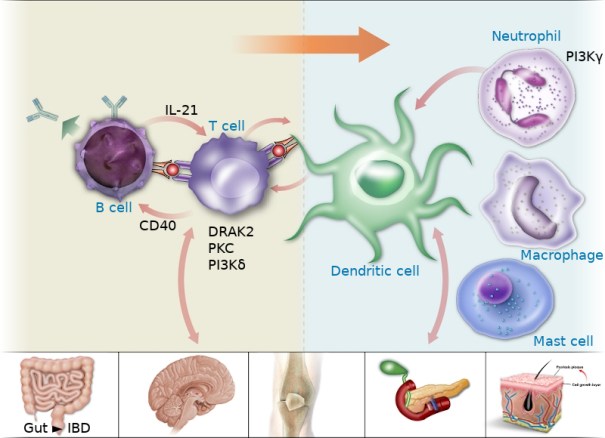
Figure 2. Graphical illustration of key immunological cell types. This forms the basis of providing a graphical-based navigation tool for GtoImmuPdb. Image copyrighted
Until now we haven’t developed the existing advanced search to cover GtoImmuPdb data types – this will be addressed in beta v3.0. We are also planning to provide video help tutorials to guide users in navigating the main GtoImmuPdb data types.
This project is supported by a 3-year grant awarded to Professor Jamie Davies at the University of Edinburgh by the Wellcome Trust (WT).


[…] Guide to IMMUNOPHARMACOLOGY (GtoImmuPdb) has issued the beta v2.2 release (see more details in the November technical blog) which includes an extension to the target advanced search. This now enables searching across the […]